 |
pyLM
v1.1
A Python Problem Solving Environment for the simulation of stochastic biological systems
|
 |
pyLM
v1.1
A Python Problem Solving Environment for the simulation of stochastic biological systems
|
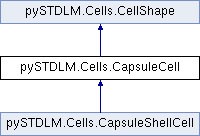
A representation for a spherical cell. More...
Public Member Functions | |
| def | __init__ |
| def | internalModelCell |
| def | computeVolume |
| def | internalAddToSim |
| def | internalTranslateCell |
 Public Member Functions inherited from pySTDLM.Cells.CellShape Public Member Functions inherited from pySTDLM.Cells.CellShape | |
| def | __init__ |
| def | getVolume |
| Volume occupied by cell NOTE: When overriding this class, you must specify a "computeVolume()" function of no arguments. More... | |
| def | boundingBox |
| Return a bounding box for the cell TODO: Make it return an orientation as well. More... | |
| def | addToSimulation |
| Add the cell to the simulation. More... | |
| def | translateCell |
| Shift a cell in space by the specified amount. More... | |
| def | createModelCell |
| Create model cell. More... | |
| def | setRegions |
| Set regions. More... | |
Public Attributes | |
| name | |
| p1 | |
| p2 | |
| radius | |
| length | |
| mins | |
| maxs | |
| membraneThickness | |
 Public Attributes inherited from pySTDLM.Cells.CellShape Public Attributes inherited from pySTDLM.Cells.CellShape | |
| name | |
| volume | |
| mins | |
| maxs | |
| membraneThickness | |
| memtype | |
| cyttype | |
A representation for a spherical cell.
This cell requires an attribute dictionary that includes "radius", "length", and "membraneThickness"
| def pySTDLM.Cells.CapsuleCell.__init__ | ( | self, | |
p1 = 3*[0.], |
|||
p2 = 3*[0.], |
|||
radius = 0., |
|||
length = 0. |
|||
| ) |
| def pySTDLM.Cells.CapsuleCell.computeVolume | ( | self | ) |
| def pySTDLM.Cells.CapsuleCell.internalAddToSim | ( | self, | |
| sim | |||
| ) |
| def pySTDLM.Cells.CapsuleCell.internalModelCell | ( | self, | |
| attr | |||
| ) |
| def pySTDLM.Cells.CapsuleCell.internalTranslateCell | ( | self, | |
| point | |||
| ) |
| pySTDLM.Cells.CapsuleCell.length |
| pySTDLM.Cells.CapsuleCell.maxs |
| pySTDLM.Cells.CapsuleCell.membraneThickness |
| pySTDLM.Cells.CapsuleCell.mins |
| pySTDLM.Cells.CapsuleCell.name |
| pySTDLM.Cells.CapsuleCell.p1 |
| pySTDLM.Cells.CapsuleCell.p2 |
| pySTDLM.Cells.CapsuleCell.radius |
 1.8.6
1.8.6