Dynamical Network Analysis
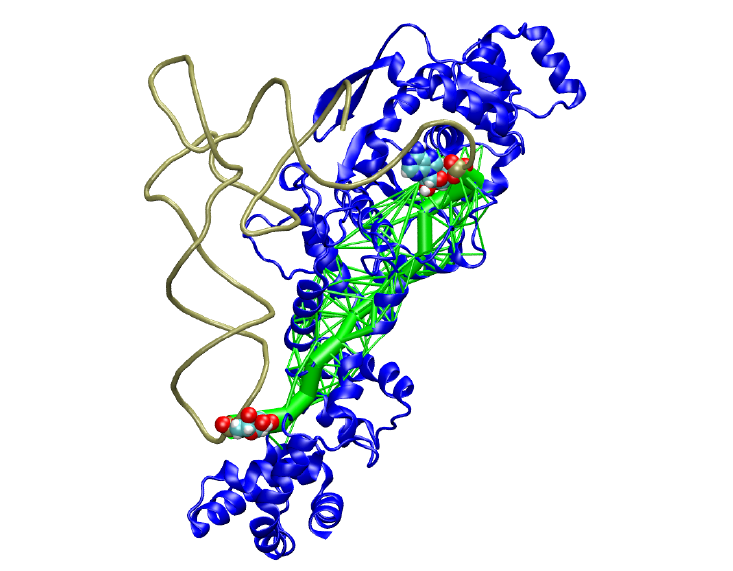
Dynamical Network Analysis is a method of characterizing allosteric signalling through biomolecular complexes.
Representative papers related to Dynamical Network Analysis:
R.W. Alexander, J. Eargle, and Z. Luthey-Schulten
Experimental and computational analysis of tRNA dynamics
FEBS Letters, 584(2):376-386, 2010
Alexis Black Pyrkosz, John Eargle, Anurag Sethi, and Zaida Luthey-Schulten
Exit strategies for charged tRNA from GluRS
J. Mol. Biol., 397:1350-1371, 2010
Anurag Sethi, John Eargle, Alexis Black, and Zaida Luthey-Schulten.
Dynamical Networks in tRNA:protein complexes
Proc. Natl. Acad. Sci. U S A, 106(16):6620-6625, 2009
Supplementary Material
Documentation
For a general introduction on how to prepare and analyze networks using NetworkView, visit the NetworkView Tutorial page.
An API reference for NetworkView has been generated as Tcldoc documentation.
Download
- Source Code: gncommunities and subopt
- LINUX x86-64 (64-bit)
- Mac OS X x86-64 (64-bit)
- Mac OS X i386 (32-bit)
- Microsoft Windows (32-bit)
Funding
This work was supported by the National Center for Supercomputing Applications [MCA03T027], the National Science Foundation [MCB-0844670] and through the Center for the Physics of Living Cells [PHY-0822613], and by the National Institutes of Health through the Center for Macromolecular Modeling and Bioinformatics [NIH-RR005969].






